20-Aug-2019
Currently there is no treatment for the deadly Hantaan infection, but new information about the virus structure could allow researchers to design a specific antiviral drug to target the virus. In an important step towards new therapies, Benoît Arragain and colleagues at the Instruct Centre FR2 in Grenoble have produced a high resolution, 3D picture of the Hantaan virus nucleocapsid.
The Hantaan virus genome is made up of three segments of RNA, which are protected by nucleoproteins, and together this structure is known as the nucleocapsid. Nucleoproteins are like building blocks that form around the RNA. Until now, it was not known exactly how the proteins assemble themselves to form a helix. This latest research has determined that 3.6 subunits are necessary to make one full rotation, and shows that each nucleoprotein of the nucleocapsid is made up of four domains: two lobes, two arms and two hinges.
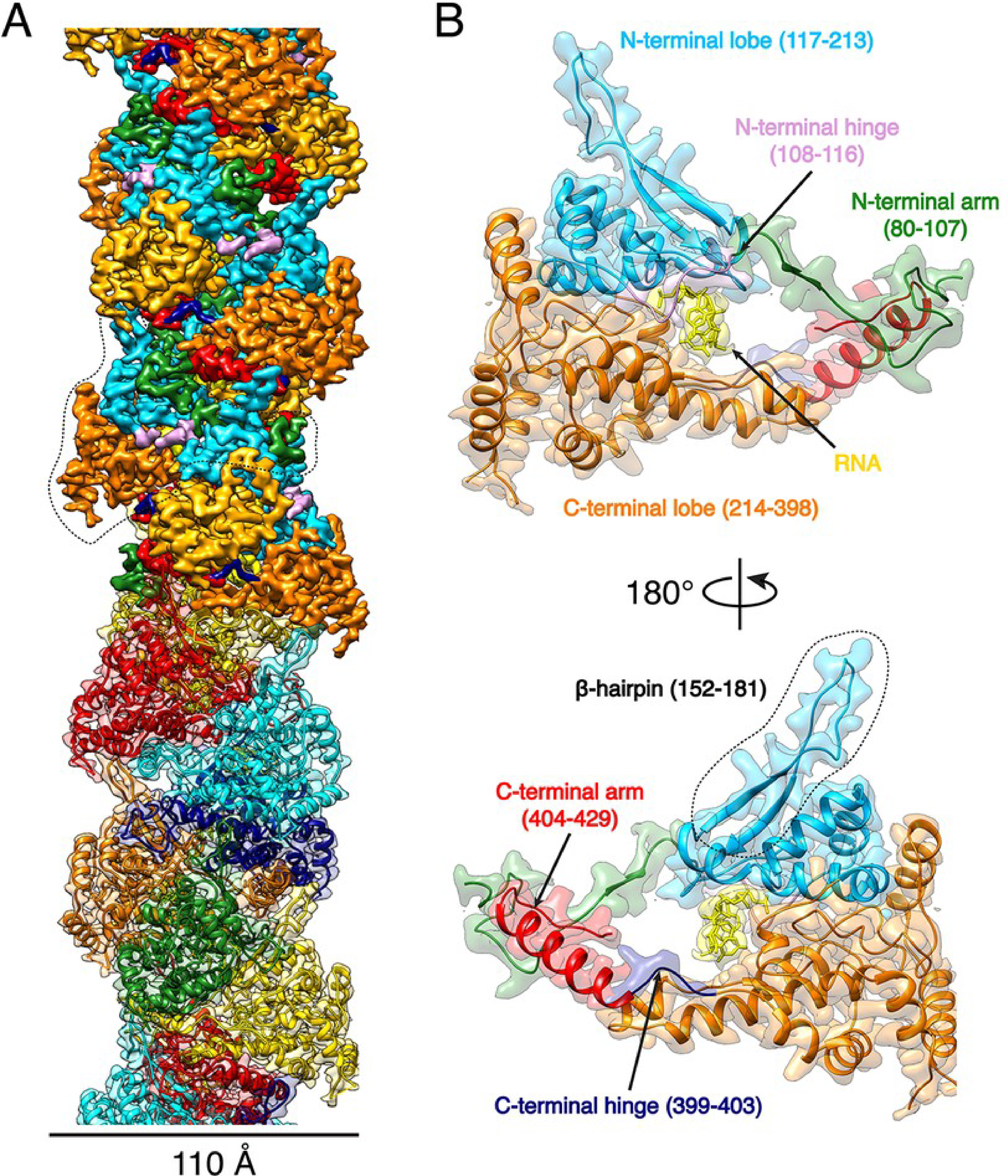
Figure 1 A and B from the paper: The helical structure of the Hantaan nucleocapsid and the six subunits that make up each repeating unit of the nucleoprotein.
Cryo-EM has allowed researchers to see, in detail, how the viral RNA and proteins interact to build the nucleocapsid. The high resolution structure (3.3 A) of the virus nucleocapsid allowed to trace the amino-acid chain into the EM density and revealed a positively-charged groove in which the RNA sits. Usually, nucleocapsids are flexible and therefore difficult to study, but cryo-EM has pin-pointed the structure of the Hantaan virus nucleocapsid with near-atomic precision. Each nucleoprotein interacts with 6 others within the nucleocapsid, which explains its rigidity.
Find out more about Cryo-EM at Instruct here.
Read more: Arragain, B., et al. (2019) High resolution cryo-EM structure of the helical RNA-bound Hantaan virus nucleocapsid reveals its assembly mechanisms. eLife 2019;8:e43075 DOI: 10.7554/eLife.43075