12-Mar-2021
Antibodies are crucial to immune protection against SARS-CoV-2, the virus responsible for the global COVID-19 pandemic. Understanding the way in which they interact with the virus, binding and neutralising, can pay significant dividends in the ongoing process of developing effective vaccines.
Researchers from the Diamond Light Source and STRUBI Instruct Centre UK collaborated to study the receptor binding sites of human antibodies to SARS-CoV-2.
Human antibodies are produced in response to infection. By isolating these from serum, monoclonal antibodies (mAbs) which are most effective at restricting infection can be generated and used to explore structural elements of the pathogen proteins. This study examined antibodies identified from recovered SARS-CoV-2 infected patients.
The study identified 377 mAbs that recognised the spike protein of the virus which binds to a cell to initiate infection via the cellular receptor ACE2. Work then focused primarily on 80 mAbs which bound directly to the virus receptor binding domain (RBD), in a variety of locations. The mAbs had varying degrees of potency at inhibiting the virus, and 19 of the 20 most potent bound to a very specific region of the cluster – blocking receptor interaction. The structure of the spike protein has been likened to a torso, with binding regions identified using anatomic features.
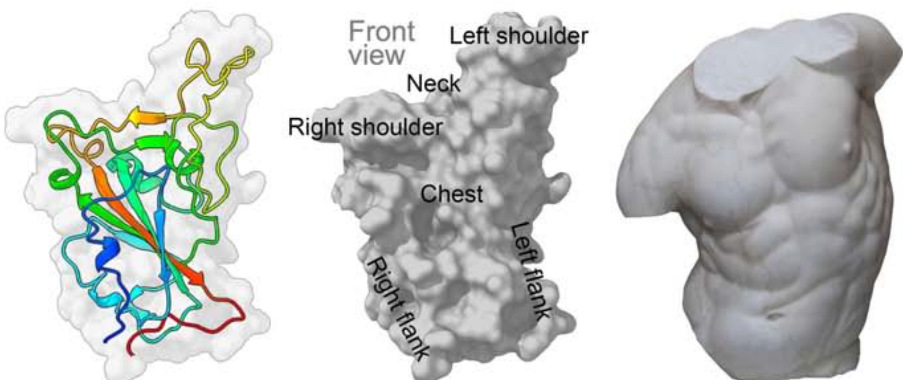
Figure 1. Pale grey RBD surface with cartoon depiction of one monomer rainbow coloured from blue (NJournal Pre-proof 29 terminus) to red (C-terminus). Also, grey surface depiction of RBD labelled to correspond to the adjacent torso - used by analogy to enable definition of epitopes.
The most potent of these mAbs were identified to bind the neck region of the spike protein, and were subsequently tested for viral inhibitory activity in vivo using transgenic mice. Tests found a significant reduction in SARS-CoV-2 induced symptoms, including weight loss, as well as reduced viral presence in the lungs and respiratory tracts. The presence of viral RNA was significantly reduced in these treated animals, compared to those treated with a control.
Administration of the most potent mAbs produced substantial benefitswhen administered one day after infection. Furthermore, even greater reduction could be achieved when utilising multiple mAbs, rather than a single clone.
This identification of binding domains and mAb potency helps advance the vaccine development program against COVID-19 by identifying key viral structural elements that can be used as therapeutic and vaccine target proteins. Access to X-ray crystallography and cryo-electron microscopy (cryo-EM) technology has enhanced the process of identifying the binding sites of the 19 potent mAbs. These technologies, available through Instruct-ERIC Centres, have enabled work to continue using remote access methods despite mobility restrictions being in place, and have contributed to the rapid developments in anti-COVID-19 responses.
Read the full text of the article: Dejnirattisai, W., Zhou D., Ginn, H. et al. The antigenic anatomy of SARS-CoV-2 receptor binding domain. Cell (2021). https://doi.org/10.1016/j.cell.2021.02.032